
History of the Arabidopsis T-DNA lines from
INRA-Versailles
Nicole Bechtold and George Pelletier
Station de Genetique et d'Amelioration des plantes,
INRA,
78026 Versailles cedex,
France
SEE NASC Update FOR RELEASE DETAILS
Origin
The deposited 148 bulks corresponding to 1480 lines came from a collection
initiated in 1992 by the discovery of a new transformation method by
vacuum infiltration (Bechtold et al.1993). In 3 years we selected 16 000 independent
transformants and screened 6,000 of them in vitro and in the greenhouse
for mutations affecting hypocotyl elongation or gametogenesis and for
flower specific promoter trapping.
Transformation method
For the in planta transformation method by vacuum infiltration we used
the Arabidopsis ecotype Wassilevskija and the binary vector pGKB5,
constructed by D. Bouchez (Bouchez et al. 1993). This vector contains a promoterless GUS
reporter gene fused to the right border, and the genes conferring kanamycin
and Basta resistance as plant selection markers.
Four-week old plants, well developed, with the first fruits formed and
secondary inflorescences appearing, were infiltrated for 20 minutes with an
Agrobacterium culture (OD600nm= 0.8) resuspended in one-third of the
initial volume in infiltration medium. Plants were then transferred to soil
and 4 weeks later the seeds were harvested. The T1 transformants were then
selected on sand sub-irrigated with water containing Basta (5-10 mg/l
phosphinothricin), transferred to soil, and grown in plastic tubes to prevent
seed contamination. The T2 seeds were harvested 6 weeks later.
pGKB5
A new plant transformation vector (see Figure 1 below) was designed, that could be
used for T-DNA tagging in Arabidopsis . The T-DNA region of this plasmid
is flanked by fragments containing the right and left borders of the TR-DNA
of pRiA4 (Jouanin et al. 1989). The GUS-nos3' reporter cassette from pBI101.1 (Jefferson et al. 1987) is inserted 40
bp away from the right border. As the GUS gene possesses its own ATG
initiation codon, it is able to produce active transcriptional or translational
gene fusions upon its insertion in the genome. The T-DNA also contains
two plant selection markers derived from pGSFR280 (De Blok et al. 1987) that confer
resistance of plant cells to kanamycin and to the herbicide Basta
(phosphinothricin).
This binary plasmid derives from a pBGS plasmid (Spratt et al. 1986), harbours a bacterial
kanamycin resistance gene, and is able to replicate both in Escherichia coli
and in Agrobacterium. The origin of replication of pRiA4, cloned as a large
8 kb BamHI fragment from pLJbB11 (Jouanin et al. 1985), confers a very high stability in
Agrobacterium under non selective conditions : Agrobacterium strains
containing pGKB5 show no detectable loss of the plasmid after 25
generations in medium lacking kanamycin. The binary vector was
introduced into several Agrobacterium disarmed strains by electroporation :
C58C1 (pMP90) (Koncz and Schell 1986), C58C1 (pGV2260) (Deblaere et al. 1985), LBA4404 (Hoekema et al. 1983) to give respectively the
strains MP5-1, GV5-2, LB5-1.
Strain MP5-1 was used for obtaining all the T-DNA lines generated in this
collection.
Figure 1. Functional map of the binary vector pGKB5.
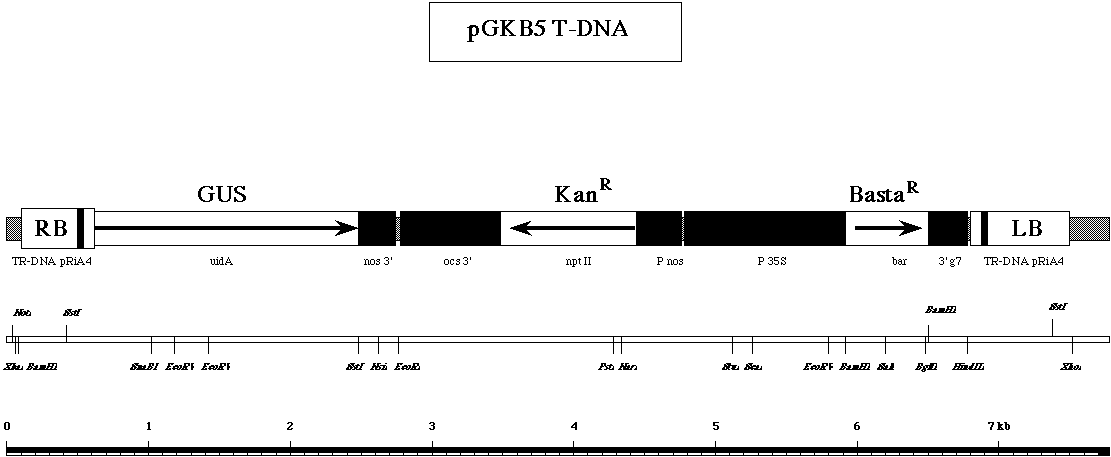
Arrows indicate coding sequences,
and black boxes promoter / terminator regions. The right and left border fragments derived
from the TR-DNA of pRiA4 from Agrobacterium rhizogenes strain A4 [3] ; the black strips
correspond to the 24 bp border sequences that serve as signals for T-DNA transfer. The
chimeric kanamycin and Basta resistance genes originate from pGSFR280 (De Blok
et al (1987). Sites for
EcoRV are indicated above the T-DNA. The hatched area corresponds to the probe used in
hybridization experiments. A large origin of replication from pRiA4 (ori pRiA4) (Jouanin et al 1985)has
been used to insure a good stability of the vector in Agrobacterium even without selection
pressure.
uidA : coding region of the beta-glucuronidase from E. coli ; nos 3' : 3' region of the nopaline
synthase gene from pTiC58 ; ocs 3' : 3' region of the octopine synthase gene from pTiAch5 ;
nptII : neomycin phosphotransferase II ; P nos : promoter region of the nopaline synthase
gene ; P 35S : promoter of the 35S transcript of the Cauliflower Mosaic Virus ; bar : coding
sequence of the basta resistance gene from Streptomyces hygroscopus ; 3' g7 : 3' region of the
gene 7 from the T-DNA of pTi15955.
Select here to view the complete sequence of the T-DNA.
Mutation screening
The T2 families for which enough seeds was produced, were screened in
vitro on medium as described by (Estelle and Somerville 1987) and modified
according to (Santoni et al. 1994).
About 100 seeds were sowed on medium containing 100 mg/l kanamycin to
estimate the number of T-DNA insertions and screen for gametophytic
mutations. The screening of the hypocotyl development mutations was
carried out without selection in light and darkness 10 days after sowing.
About 50 seeds of each T2 families were also sowed on compost without
selection in the greenhouse to screen early development and gametogenesis
mutations.
GUS screening
The GUS activity of 2,000 first T2 lines was tested histochemically on 15-
days-old seedlings grown in vitro either under light or dark conditions (Mollier et al. 1995).
For all the families we tested the GUS activity on flowers and siliques taken
from 6 weeks old plants grown in the greenhouse.
Seed production
Because of the low number of T2 seeds it was necessary to multiply them
and send the T3 generation to the Nottingham Arabidopsis Stock Centre. T3 seeds come from the bulk
of about 50 T2 plants for each family. Seeds were stored at 4C and 15 % RH.
The aliquots contain hemizygous, homozygous and wild type seeds.
Available data
The available data were recorded through analysis of T2 seedlings in vitro
and flowering plants in the greenhouse. This information includes:
- in vitro segregation on about 100 seeds sowed on kanamycin 100
mg/l (corresponding to 3:1 segregation, 15:1, 63:1, 2:1, ?= number of
resistant plants 3.86
for all the possible segregations).
- GUS expression at seedling stage in vitro for a limited number of
lines [3].
- GUS expression in flowers and siliques in the greenhouse.
- seedling observations in vitro
- plant observations in the greenhouse.
Unavailable lines
A number of lines are not yet available through NASC because we or our
collaborators were interested in:
- 1:1 segregation for kanamycin resistance.
- lines containing about 1/4 sterile or partial sterile mutants.
- hypocotyl development mutants.
- leaf morphology mutants.
- leaf necrosis mutants.
- germination mutants.
- yellow-green mutants.
- specific GUS expression in flower and siliques.
- some specific GUS expression in leaves.
- lines we didn't test in vitro because they yielded only a few T2 seeds
(1/3 of the lines, but they will be tested in T3 generation).
Restriction for the use of the Versailles-INRA T-DNA lines
These lines are freely distributed by the NASC for fundamental research
purposes. Commercial application of the information obtained from the
study of these lines is not possible without prior agreement with INRA.
Important notes
- The segregation of the kanamycin marker was estimated on about 100
seedlings. This is not always enough to distinguish between segregations
like 2:1 and 3:1 for example. It is then necessary to verify the segregation
with more seeds (about 300).
- For several lines no resistant progeny were recovered on kanamycin, but
in Southern analysis some of them do contain the T-DNA. We decided to
send all the lines even if all the plants were kanamycin sensitive. These
lines were noted "?" in the segregation data.
- We also have observed differences between GUS expression at seedling
stage in vitro and in the greenhouse, and some unrepeatable GUS
expression in flowers and siliques (GUS expression varied with greenhouse
conditions).
- Not all the GUS expression observed was necessarily tissue-specific. We
just tested the specificity of GUS expression in the flowers and siliques.
- The lines were not always grown and harvested in the same climatic
conditions, and it is possible that this lead to some differences in the
germination rate.
- In this collection, we observed that 56 % of T-DNA integration are at a
single locus, and by Southern 75 % of them are in tandem (Bechtold et al. 1993).
- From the first studies of mutants provided by this collection, it seems that
about 1/4 of the lines showing a segregation of mutants/ wild type near to
1:3 will be tagged. As already described by Feldmann, the in planta methods
also appear to have a mutagenic effect.
References
N Bechtold , J Ellis, G Pelletier. (1993) In planta Agrobacterium mediated gene
transfer by infiltration of adult Arabidopsis thaliana plants. C. R. Acad. Sci.
Paris, Sciences de la vie/Life Sciences 316, 1194-9
D. Bouchez, C. Camilleri, M. Caboche.(1993) A binary vector based on Basta
resistance for in planta transformation of Arabidopsis thaliana C. R. Acad.
Sci. Paris, Sciences de la vie/Life sciences 316, 1188-93
Jouanin L., Bouchez D., Drong R. F., Tepfer D., Slightom J. L. (1989)
Analysis of TR-DNA / plant junctions in the genome of a Convolvulus
arvensis clone transformed by Agrobacterium rhizogenes strain A4. Plant
Mol. Biol. 12, 75-85.
Jefferson R. A., Kavanagh T. A., Bevan M. W.( 1987) GUS fusions , beta-
glucuronidase as a sensitive and versatile gene fusion marker in higher
plants. EMBO J. 6, 3901-3907.
De Blok M., Botterman J., Vanderwiele M., Dockx J., Thoen C., Gossele V.,
Rao Movva N., Thompson C., Van Montagu M., Leemans J. (1987)
Engineering herbicide resistance in plants by expression of a detoxifying
enzyme.EMBO J. 6, 2513-2518.
Spratt B. G., Hedge P. J., te Heesen S., Edelman A., Broome-Smith J. K.
(1986) Kanamycin-resistant vectors that are analogs of plasmids pUC8, pUC9,
pEMBL8 and pEMBL9. Gene 41, 337-342.
Jouanin L., Vilaine F., d'Enfert C., Casse-Delbart F. (1985) Localization and
restriction maps of the replication origin regions of the plasmids of
Agrobacterium rhizogenes strain A4. Mol. Gen. Genet. 210, 370-374.
Koncz C., Schell J.( 1986) The promoter of TL-DNA gene 5 controls the
tissue-specific expression of chimaeric genes carried by a novel type of
Agrobacterium binary vector. Mol. Gen. Genet. 204, 383-396.
Deblaere R., Bytebier B., De Greve H., Deboeck F., Schell J., Van Montagu
M., Leemans J. (1985) Efficient octopine Ti plasmid vectors for
Agrobacterium-mediated gene transfer to plants. Nucl. Acids Res. 13, 4777-
4788.
Hoekema A., Hirsch P. R., Hooykas P. J. J., Schilperoort R. A. (1983). A
binary plant vector strategy based on separation of vir- and T-region of the
Agrobacterium tumefaciens Ti plasmid. Nature 303, 179-180.
Estelle M.A. and Somerville C.R. (1987) Auxin-resistant mutants of
Arabidopsis thaliana with an altered morphology. Mol. Gen. Genet. 206, 200-6
Santoni V., Bellini C., Caboche M. (1994) Use of two-dimensional protein-
pattern analysis for the characterization of Arabidopsis thaliana mutants.
Planta 192, 557-66
P. Mollier, P. Montoro, M. Delarue, N. Bechtold, C. Bellini, G. Pelletier.
Promoterless gusA expression in a large number of Arabidopsis thaliana (1995)
transformants obtained by the in planta infiltration method. C.R.Acad.Sci.
Paris, Sciences de la vie/Life sciences 318, 465-74

